ExPSIBLAST answer and ExPSIBLAST: Difference between pages
(Created page with "Note: E-values etc. are found November 8, 2023. == When BLAST fails == * '''QUESTION 1''': How many significant hits does BLAST find (E-value < 0.005)? Answer: No sequences with E-value below 0.005. ==Trying another approach== * '''QUESTION 2''': How many significant hits does BLAST find (E-value < 0.005)? Answer: After the first iteration, 363 hits are found. * '''QUESTION 3''': How large a fraction of the query sequence do the significant hits match (excluding the...") |
|||
| Line 1: | Line 1: | ||
Originally written by: Morten Nielsen — some editing by Rasmus Wernersson and Bent Petersen — new version by Henrik Nielsen. | |||
== When BLAST fails == | ==Introduction== | ||
Earlier in the course you have used the BLAST program to perform fast alignments of DNA and protein sequences. As shown in today's lecture, BLAST will often fail to recognize relationships between proteins with low sequence similarity. In today's exercise, you will use the iterative BLAST program (PSI-BLAST) to calculate sequence profiles and see how such profiles can be used to: | |||
* Identify relationships between proteins with low sequence similarity | |||
* Identify conserved residues in protein sequences (residues important for the structural stability or function of the protein) | |||
===Links=== | |||
* NCBI BLAST: http://www.ncbi.nlm.nih.gov/BLAST/ | |||
<!-- * [https://services.healthtech.dtu.dk/service.php?Blast2logo Blast2logo] is a tool for visualization of protein sequence profiles and identification of conserved residues. | |||
--> | |||
==When BLAST fails== | |||
Say you have a sequence [https://teaching.healthtech.dtu.dk/material/22111/files/Query1.txt Query] (pasted below) and you want to make predictions about its function and structure. As seen earlier in the course, you will most often use BLAST to do this. However what happens when BLAST fails? | |||
>QUERY1 | |||
MKDTDLSTLLSIIRLTELKESKRNALLSLIFQLSVAYFIALVIVSRFVRYVNYITYNNLV | |||
EFIIVLSLIMLIIVTDIFIKKYISKFSNILLETLNLKINSDNNFRREIINASKNHNDKNK | |||
LYDLINKTFEKDNIEIKQLGLFIISSVINNFAYIILLSIGFILLNEVYSNLFSSRYTTIS | |||
IFTLIVSYMLFIRNKIISSEEEEQIEYEKVATSYISSLINRILNTKFTENTTTIGQDKQL | |||
YDSFKTPKIQYGAKVPVKLEEIKEVAKNIEHIPSKAYFVLLAESGLRPGELLNVSIENID | |||
LKARIIWINKETQTKRAYFSFFSRKTAEFLEKVYLPAREEFIRANEKNIAKLAAANENQE | |||
IDLEKWKAKLFPYKDDVLRRKIYEAMDRALGKRFELYALRRHFATYMQLKKVPPLAINIL | |||
QGRVGPNEFRILKENYTVFTIEDLRKLYDEAGLVVLE | |||
[File:blastp_pdb.png|center|frame|Partial screenshot of the Blast interface. The red arrow shows the settings change to the database to pdb]] | |||
Go to the [http://www.ncbi.nlm.nih.gov/BLAST BLAST] web-site at NCBI. Select <u>blastp</u> as the algorithm. Paste in the query sequence. Change the database from nr to <u>pdb</u>, and press <u>BLAST</u>. | |||
* '''QUESTION 1''': How many significant hits does BLAST find (E-value < 0.005)? | * '''QUESTION 1''': How many significant hits does BLAST find (E-value < 0.005)? | ||
==Trying another approach== | ==Trying another approach== | ||
* '''QUESTION 2''': How many significant hits does BLAST find (E-value < 0.005)? | <!-- [[File:PsiBlast_MN.png|center|frame|Partial screen shoot of the Psi-Blast interface. Here, the Protein Data Bank (pdb) has been selected as database.]] --> | ||
[[File:psiblastp_nr.png|250px|center|frame|Partial screenshot of the Psi-Blast interface. The red arrow shows the settings change to Psi-Blast.]] | |||
Now go back to the [http://www.ncbi.nlm.nih.gov/BLAST BLAST] web-site. Paste in the query sequence [https://teaching.healthtech.dtu.dk/material/22111/files/Query1.txt Query1]. This time, set the database to <u>nr</u> and select <u>PSI-BLAST (Position-Specific Iterated BLAST)</u> as the algorithm. '''IMPORTANT:''' To ease the load on the NCBI server, limit the search to Archaea (TaxID 2157) when searching Query1 in nr. | |||
* '''QUESTION 2''': How many significant hits does BLAST find (E-value < 0.005)? ('''Tip:''' you can see the number by selecting all significant hits (clicking <u>All</u> under <u>Sequences producing significant alignments with E-value BETTER than threshold</u>) and then looking at the number of selected hits) | |||
* '''QUESTION 3''': How large a fraction (Query coverage) of the query sequence do the significant hits match (excluding the identical match)? | |||
* '''QUESTION 4''': Do you find any PDB hits among the significant hits? ('''Tip:''' look for a PDB identifier in the <u>Accession</u> column — a PDB identifier is a 4 character code, where the first character is a number, followed by a single letter chain name, such as "1XYZ_A") | |||
===Constructing the PSSM=== | |||
<div style="background-color: lightyellow; border: solid thin grey;"> | |||
:'''Note:''' If you see the error message “<u>Entrez Query: txid2157 [ORGN] is not supported</u>”, then click <u>Recent Results</u> in the upper right part of the BLAST window, select your most recent search, and try again. | |||
</div> | |||
Now run a second BLAST iteration in order to construct a PSSM (Position-Specific Scoring Matrix). Press the <u>Run</u> button at <u>Run PSI-Blast iteration 2</u> (you can find it at both the bottom and top of the results table). | |||
* '''QUESTION 5''': How many significant hits does BLAST find (E-value < 0.005)? | * '''QUESTION 5''': How many significant hits does BLAST find (E-value < 0.005)? | ||
* '''QUESTION 6''': How large a fraction of the query sequence do the 20 most significant hits match (do not include the first hit since this is identical to the query)? | |||
* '''QUESTION 7''': Why does BLAST come up with more significant hits in the second iteration? Make sure you answer this question and understand what is going on! | |||
===Saving and reusing the PSSM=== | |||
This time, we will not ask you to look for PDB identifiers manually among the significant hits. Instead, you should save the PSSM that PSI-BLAST has created and use it for searching PDB directly. | |||
Go to the top of the PSI-BLAST output page and click <u>Download All</u>, then click <u>PSSM</u>. Save the file to a place on your computer where you can find it again! You can take a look at this file using Geany, but it is really not meant to be human-readable. | |||
Then, open ''a new BLAST window'' (this is important—you need your first BLAST window again later) where you again select PSI-BLAST as the algorithm. Select <u>pdb</u> as the database. Do ''not'' limit your search to Archaea this time. Click on <u>Algorithm parameters</u> to show the extended settings. Click the button next to <u>Upload PSSM</u> and select the file you just saved. '''Note:''' You don't have to paste the query sequence again, it is stored in the PSSM! | |||
* '''QUESTION 8''': Do you find any significant PDB hits now? If yes, how many? | * '''QUESTION 8''': Do you find any significant PDB hits now? If yes, how many? | ||
* '''QUESTION 9''': What are the PDB identifiers and the E-values for the two best PDB hits? | * '''QUESTION 9''': What are the PDB identifiers and the E-values for the two best PDB hits? | ||
* '''QUESTION 10''': What are the values for Query coverage, sequence identity, and sequence similarity (Positives) for the two best PDB hits? ('''Tip:''' click on the description to get to the actual alignment between the query sequence and the PDB hit)? | |||
* '''QUESTION 11''': What is the function of these proteins? | |||
* '''QUESTION | ===One more round=== | ||
Let's try one more iteration of PSI-BLAST: | |||
* Go back to your first BLAST window (the one with the results from the <u>nr</u> database limited to Archaea) and press the <u>Run</u> button at <u>Run PSI-Blast iteration 3</u>. | |||
* Save the resulting PSSM file (make sure you give it a different name!). | |||
* Launch a new PSI-BLAST search against <u>pdb</u> in all organisms using this PSSM (you may have to click on <u>Clear</u> to erase your first PSSM file from the server). | |||
* '''QUESTION 12''': Answer questions 8-10 again for the new search. | |||
''' | ==Finding a remote homolog (on your own)== | ||
PSI-Blast is not only useful for finding a remote homolog in a specific database such as PDB — now it is time to search the broader database "Reference proteins" (<u>refseq_protein</u>). ('''Note:''' we would have liked to do this exercise in the broadest database <u>nr</u>, but that search runs into technical problems). PSI-BLAST can be used in the same way for finding a remote homolog in a specific organism or taxonomic group. Your task in this round is to find out whether the protein with the UniProt ID '''GPAA1_HUMAN''' has a homolog in the genus ''Trypanosoma'' (unicellular parasites which cause diseases like sleeping sickness or Chaga's disease). | |||
* First, try a standard BlastP (where you set <u>Organism</u> to ''Trypanosoma'', <u>Database</u> to <u>refseq_protein</u> ('''not''' refseq_select), switch the <u>Low complexity regions</u> filter off, and set the E-value threshold to 10). | |||
* '''QUESTION 13''': Do you find any significant (E<0.005) hits? What is the E-value of the best hit? | |||
* Then, try PSI-BLAST. '''Hint:''' You need to search in all organisms (still using refseq_protein) to build a PSSM, then save your PSSM and use that to search in ''Trypanosoma''. | |||
* '''QUESTION 14''': How many significant (E<0.005) hits do you find now? What is the E-value of the best hit? | |||
<!-- | |||
==Identifying conserved residues== | |||
[[File: Logo.png|right|frame|thumb|Logo of a sequence profile spanning residues 279-296. The logo is calculated from a Psi-Blast profile]] | |||
We now return to the Query sequence you used in questions 1-12. You have now (hopefully) identified a structural relationship between the Query sequence and a protein sequence in the PDB database of protein structures. Say you would like to validate this relationship. This could be done by mutating (substituting) essential residues in the query sequence and test if the protein function (or structure) is affected by these mutations. | |||
The protein sequence of the query is large (more than 400 amino acids) and a complete mutation study including all residues would be extremely costly. Instead one can use PSI-BLAST and sequence profiles to identify conserved residues that are likely to be essential for the protein structure and/or protein function. | |||
Below you find a set of 8 residues from the Query protein sequence. You shall use the PSI-BLAST and Blast2logo programs to select four of the eight residues for a mutagenesis study (you shall select the four residues based on sequence conservation only). | |||
* (a): H271 | |||
* (b): R287 | |||
* (c): E290 | |||
* (d): Y334 | |||
* (e): F371 | |||
* (f): R379 | |||
* (g): R400 | |||
* (h): Y436 | |||
You shall use the [https://services.healthtech.dtu.dk/service.php?Blast2logo Blast2logo] server to identify which residues are conserved in the [https://teaching.healthtech.dtu.dk/material/36611/files/Query1.txt Query] protein sequence. Go to the Blast2logo server and upload the Query sequence. Set the Blast database to <u>NR70</u>, set the logo type to <u>Shannon</u> and press submit (note it might take some (10-15) minutes before your job is completed). If the job does not complete, or if you don't have the patience to wait, you can find the output following this link [https://teaching.healthtech.dtu.dk/material/36611/files/Blast2logo_Query1_frame.htm Blast2logo output]. | |||
' | |||
When the job is completed you should see the logo-plot on the website. You can download it in PDF format. To improve the readability of the logo, you can also click on the <u>Customize visualization using Seq2Logo</u> button. In doing this your are transferred to the Seq2Logo web server. Here, just leave all options as default, and press submit. | |||
* '''QUESTION 15''': Spend a little time looking at the logo plot. Can you understand why the logo is so flat for the first 150 residues (how large a fraction of the query section did the Blast search cover)? | |||
* '''QUESTION 16''': Which four of the eight residues listed above are most conserved and hence most likely to be essential for the protein stability and/or function? | |||
--> | |||
<!-- | |||
== | ===Homology modelling === | ||
You shall use the [http://www.sbg.bio.ic.ac.uk/phyre2/ Phyre2] program to validate if the structural properties of the four most conserved residues from question Q12 indeed could form an active site. Phyre is a program for protein homology modelling. This program takes as input a protein amino acids sequence and produces as output a 3D protein structure based on single template homology modelling. Go to the Phyre web-site and upload the [https://teaching.healthtech.dtu.dk/material/36611/files/Query1.txt Query] sequence. Note it might take some (10-20) minutes before your job is completed. To save you time, I have run the calculation for you. You can find the output here [http://www.sbg.bio.ic.ac.uk/servers/phyre/qphyre_scripts/results.cgi?jobid=070ac42bdea13d4e Phyre output]. | |||
Find the PDB hit identified by PSI-BLAST (you can click on the on the 3D model of the protein to get the relevant PDB filel). | |||
You shall use the [https://services.healthtech.dtu.dk/service.php?CPHmodels CPHmodels] server to validate if the structural properties of the four most conserved residues from question Q14 indeed could form an active site. CHPmodels is a program for protein homology modelling. This program takes as input a protein amino acids sequence and produces as output a 3D protein structure based on single template homology modelling. Go to the CPHmodels web-site and upload the [https://teaching.healthtech.dtu.dk/material/36611/files/Query1.txt Query] sequence. Note it might take some (10-20) minutes before your job is completed. To save you time, I have run the calculation for you. You can find the output here: [http://www.cbs.dtu.dk/services/CPHmodels/teaching/query1.html CPHmodels output] | |||
The output from CPHmodels is not straightforward to interpret. However, the method provides a Z-score in situations where the query and template shares little sequence similarity. As a rule of thumb, a Z-score greater than 10 will signify a reliable model. You find the template used by CPHmodels and the Z-score for the model in the last part of the output file. | |||
* '''QUESTION 17''': Does CPHmodels agree that the hit identified by PSI-Blast hit is significant? | * '''QUESTION 17''': Does CPHmodels agree that the hit identified by PSI-Blast hit is significant? | ||
Download the homology model made by CPHmodels (click on the query.pdb link), and open the model file in Pymol. If you do not have Pymol installed on your computer, you can find it on Campusnet in the Pymol folder. Show the location of the four essential residues from question Q14 on the structure. | |||
* '''QUESTION 18''': Could the residues form an active site? | * '''QUESTION 18''': Could the residues form an active site? | ||
--> | |||
==Concluding remarks== | |||
Now you have seen the power of sequence profiles in general and the PSI-BLAST program in particular. Using sequence profiles you have been able to identify a relationship between protein sequences far below 30% sequence similarity. Further, you have made qualified predictions on the protein function and selected a set of essential amino acids suitable for experimental validation of the structural and functional predictions. | |||
Revision as of 10:30, 5 November 2025
Originally written by: Morten Nielsen — some editing by Rasmus Wernersson and Bent Petersen — new version by Henrik Nielsen.
Introduction
Earlier in the course you have used the BLAST program to perform fast alignments of DNA and protein sequences. As shown in today's lecture, BLAST will often fail to recognize relationships between proteins with low sequence similarity. In today's exercise, you will use the iterative BLAST program (PSI-BLAST) to calculate sequence profiles and see how such profiles can be used to:
- Identify relationships between proteins with low sequence similarity
- Identify conserved residues in protein sequences (residues important for the structural stability or function of the protein)
Links
- NCBI BLAST: http://www.ncbi.nlm.nih.gov/BLAST/
When BLAST fails
Say you have a sequence Query (pasted below) and you want to make predictions about its function and structure. As seen earlier in the course, you will most often use BLAST to do this. However what happens when BLAST fails?
>QUERY1 MKDTDLSTLLSIIRLTELKESKRNALLSLIFQLSVAYFIALVIVSRFVRYVNYITYNNLV EFIIVLSLIMLIIVTDIFIKKYISKFSNILLETLNLKINSDNNFRREIINASKNHNDKNK LYDLINKTFEKDNIEIKQLGLFIISSVINNFAYIILLSIGFILLNEVYSNLFSSRYTTIS IFTLIVSYMLFIRNKIISSEEEEQIEYEKVATSYISSLINRILNTKFTENTTTIGQDKQL YDSFKTPKIQYGAKVPVKLEEIKEVAKNIEHIPSKAYFVLLAESGLRPGELLNVSIENID LKARIIWINKETQTKRAYFSFFSRKTAEFLEKVYLPAREEFIRANEKNIAKLAAANENQE IDLEKWKAKLFPYKDDVLRRKIYEAMDRALGKRFELYALRRHFATYMQLKKVPPLAINIL QGRVGPNEFRILKENYTVFTIEDLRKLYDEAGLVVLE
[File:blastp_pdb.png|center|frame|Partial screenshot of the Blast interface. The red arrow shows the settings change to the database to pdb]]
Go to the BLAST web-site at NCBI. Select blastp as the algorithm. Paste in the query sequence. Change the database from nr to pdb, and press BLAST.
- QUESTION 1: How many significant hits does BLAST find (E-value < 0.005)?
Trying another approach
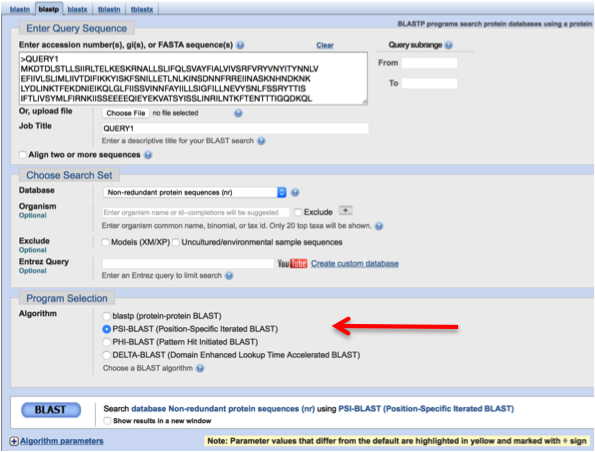
Now go back to the BLAST web-site. Paste in the query sequence Query1. This time, set the database to nr and select PSI-BLAST (Position-Specific Iterated BLAST) as the algorithm. IMPORTANT: To ease the load on the NCBI server, limit the search to Archaea (TaxID 2157) when searching Query1 in nr.
- QUESTION 2: How many significant hits does BLAST find (E-value < 0.005)? (Tip: you can see the number by selecting all significant hits (clicking All under Sequences producing significant alignments with E-value BETTER than threshold) and then looking at the number of selected hits)
- QUESTION 3: How large a fraction (Query coverage) of the query sequence do the significant hits match (excluding the identical match)?
- QUESTION 4: Do you find any PDB hits among the significant hits? (Tip: look for a PDB identifier in the Accession column — a PDB identifier is a 4 character code, where the first character is a number, followed by a single letter chain name, such as "1XYZ_A")
Constructing the PSSM
- Note: If you see the error message “Entrez Query: txid2157 [ORGN] is not supported”, then click Recent Results in the upper right part of the BLAST window, select your most recent search, and try again.
Now run a second BLAST iteration in order to construct a PSSM (Position-Specific Scoring Matrix). Press the Run button at Run PSI-Blast iteration 2 (you can find it at both the bottom and top of the results table).
- QUESTION 5: How many significant hits does BLAST find (E-value < 0.005)?
- QUESTION 6: How large a fraction of the query sequence do the 20 most significant hits match (do not include the first hit since this is identical to the query)?
- QUESTION 7: Why does BLAST come up with more significant hits in the second iteration? Make sure you answer this question and understand what is going on!
Saving and reusing the PSSM
This time, we will not ask you to look for PDB identifiers manually among the significant hits. Instead, you should save the PSSM that PSI-BLAST has created and use it for searching PDB directly.
Go to the top of the PSI-BLAST output page and click Download All, then click PSSM. Save the file to a place on your computer where you can find it again! You can take a look at this file using Geany, but it is really not meant to be human-readable.
Then, open a new BLAST window (this is important—you need your first BLAST window again later) where you again select PSI-BLAST as the algorithm. Select pdb as the database. Do not limit your search to Archaea this time. Click on Algorithm parameters to show the extended settings. Click the button next to Upload PSSM and select the file you just saved. Note: You don't have to paste the query sequence again, it is stored in the PSSM!
- QUESTION 8: Do you find any significant PDB hits now? If yes, how many?
- QUESTION 9: What are the PDB identifiers and the E-values for the two best PDB hits?
- QUESTION 10: What are the values for Query coverage, sequence identity, and sequence similarity (Positives) for the two best PDB hits? (Tip: click on the description to get to the actual alignment between the query sequence and the PDB hit)?
- QUESTION 11: What is the function of these proteins?
One more round
Let's try one more iteration of PSI-BLAST:
- Go back to your first BLAST window (the one with the results from the nr database limited to Archaea) and press the Run button at Run PSI-Blast iteration 3.
- Save the resulting PSSM file (make sure you give it a different name!).
- Launch a new PSI-BLAST search against pdb in all organisms using this PSSM (you may have to click on Clear to erase your first PSSM file from the server).
- QUESTION 12: Answer questions 8-10 again for the new search.
Finding a remote homolog (on your own)
PSI-Blast is not only useful for finding a remote homolog in a specific database such as PDB — now it is time to search the broader database "Reference proteins" (refseq_protein). (Note: we would have liked to do this exercise in the broadest database nr, but that search runs into technical problems). PSI-BLAST can be used in the same way for finding a remote homolog in a specific organism or taxonomic group. Your task in this round is to find out whether the protein with the UniProt ID GPAA1_HUMAN has a homolog in the genus Trypanosoma (unicellular parasites which cause diseases like sleeping sickness or Chaga's disease).
- First, try a standard BlastP (where you set Organism to Trypanosoma, Database to refseq_protein (not refseq_select), switch the Low complexity regions filter off, and set the E-value threshold to 10).
- QUESTION 13: Do you find any significant (E<0.005) hits? What is the E-value of the best hit?
- Then, try PSI-BLAST. Hint: You need to search in all organisms (still using refseq_protein) to build a PSSM, then save your PSSM and use that to search in Trypanosoma.
- QUESTION 14: How many significant (E<0.005) hits do you find now? What is the E-value of the best hit?
Concluding remarks
Now you have seen the power of sequence profiles in general and the PSI-BLAST program in particular. Using sequence profiles you have been able to identify a relationship between protein sequences far below 30% sequence similarity. Further, you have made qualified predictions on the protein function and selected a set of essential amino acids suitable for experimental validation of the structural and functional predictions.